Data & biology
Popular emphasis has been on the architectures driving progress in AI, such as attention, transformers, and diffusion models. However, this focus often overshadows a critical element: the role of data.
Data can not be taken for granted, especially in the world of biology. Unlike images and text, which can be retained at little cost in large volumes from the internet, acquiring biological data is resource intensive. Collecting biological data at scale requires large amounts of time and money. In fact, many AI-first therapeutics firms are wagering on their data-generation capabilities to gain an edge. Ideally, this strategy leads to more efficient models and faster research discoveries, creating a “flywheel” that unlocks novel biological insights. This sounds like an attractive proposition but it has not been strongly proven out in the first-generation businesses – a topic I’ve explored in a previous post.
Undoubtedly, more data is essential for improving both generative and discriminative tasks in life sciences. Whether the en vogue private strategy succeeds or not, acquiring vast data volumes is key to enhancing models. Richard Sutton’s Bitter Lesson can’t be reiterated enough:
…general methods that leverage computation are ultimately the most effective, and by a large margin…We have to learn the bitter lesson that building in how we think does not work in the long run…breakthrough progress eventually arrives by an opposing approach based on scaling computation by search and learning.1
As model architectures and training strategies continue to evolve, the primary challenge in biology becomes data acquisition. Michael Bronstein recently highlighted this impending change, anticipating that "3rd generation" biotech companies will rely on platforms designed to scale up data collection, likely at the cost of measurement resolution.
Given the resource-intensive nature of acquiring biological data, a critical question emerges: How does biological data scale in the forthcoming years? In this series of posts I will explore how the data acquisition landscape is evolving across the bioML stack of sequences, structures and systems.
My expertise lies in modeling scRNA-seq data, so I'll begin with systems. In this context, "systems" refers to concentration or abundance, reflecting the dynamic shifts in the quantity and composition of biomolecules in cells or organisms.
Self-supervised learning & compute
Let's begin by detailing how recent approaches scale computation to improve machine learning systems. The concept of self-supervised learning (SSL) is at the core of this advancement.
The anchor moment is BERT (Devlin, 2018) which introduced the masked language modeling (MLM) objective for pretraining large language models.2 The MLM objective involves training the model to predict missing words within a sentence, enabling it to learn strong statistical priors about natural language. This is the essence of SSL: leveraging unlabeled data to scale computation. The innovation here is exploiting the accessibility and low cost of unlabeled textual data. BERT showed you can pretrain a model on unlabeled data, and then use this model with strong priors to improve performance on “downstream” supervised tasks. This is a form of transfer learning, which involves two stages: (1) pretraining a model using SSL, and then (2) fine-tuning it on specific tasks with labeled datasets.
What does this allow you to do? Scale compute – please don’t forget the Bitter Lesson. We saw the pretraining paradigm flood all other information domains. In biology we have seen protein language models (ESM, Rives, 2021), genomic language models (Nucleotide Transformer, Dalla-Torre, 2023) and scRNA-seq language models (Geneformer, Theodoris, 2023).3 These are just a few early examples among many contributions, which I plan to discuss in more detail throughout this series of posts. To delve deeper, you can refer to my tweets on scRNA-seq foundation models and genomic foundation models.
scRNA-seq data trends
Most are familiar with Moore's Law, which predicts the doubling of transistor density every two years, leading to exponential advancements in technology and more affordable, scalable computing. A similar exponential trend is apparent in DNA sequencing technology – a topic I'll explore more in a future post on “sequences.”
The scaling trends in single-cell RNA sequencing (scRNA-seq) data become particularly significant when considering the role of computation in ML systems alongside the challenges of collecting biological data. I took a look at the volume of publicly available scRNA-seq data over the past decade using the CZI CELLxGENE resource.
On top we have the cumulative number of cells over time, and on bottom we have the cumulative number of tokens over time. I define a single “token” as a measured gene – this does not account for read depth. Intriguingly, these trends suggest the possibility of technology improvements surpassing a linear progression.
The red line is a linear forecast, and the green line is an exponential forecast. It is hard to know how assay technology is truly impacting measurement scales as we are in early innings. While my knowledge of evolving assay technologies is somewhat limited, it's clear that even standard assays have seen significant scaling advancements, notably through techniques like multiplexing. Large-scale data generation by major consortiums plays a role, though increased spending likely isn't the sole factor that explains the quickly increasing data volumes. A more comprehensive analysis would really examine the unit cost of scRNA-seq over time.
NLP scaling law framework
At the time of writing, the CELLxGENE resource boasts approximately 60M cells and 120B tokens. Let's anchor this to the leading applied AI research fields to add some context. NLP has the most thorough benchmarks of compute scaling laws so we can look here for a framework. Before doing so I want to highlight that there is a major risk in analogizing biological models to textual language models. The data generation processes are very different as are the modeling objectives and approaches.
That being said, I think this framework can be helpful in building rough intuition. GPT-3 was trained on 300B tokens (1 token ≈ 0.75 words) and has 175B parameters. Prior to GPT-3, OpenAI published, “Scaling Laws for Neural Language Models” in January 2020.4 They empirically determined the relationship between model loss and dataset size, and model loss and parameter count. Following up on this was Deepmind’s Chinchilla, presented in “Training Compute-Optimal Large Language Models.”5 In this work the goal was to determine the optimal model size and dataset size for a given compute budget (FLOPs). The short story is that they found large models like 175B parameter GPT-3 were significantly undertrained (not enough data:params). They use their findings to train a 70B parameter model on 1.4T tokens. Chinchilla bested other SOTA models at the time while also being significantly smaller (inference is cheaper and faster). Here is a great Less Wrong post if you’d like to dive into the details.
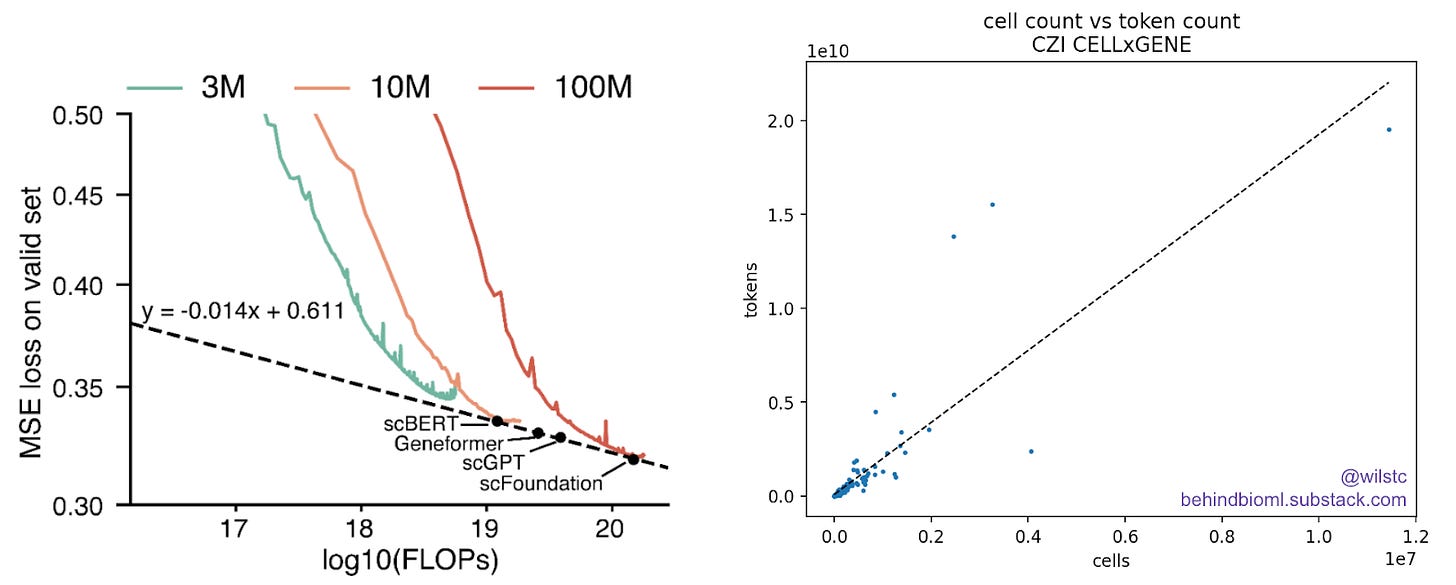
For reference, the scFoundation paper did a modest model-scaling experiment. They evaluated FLOPs versus loss at different model sizes. The 100M parameter scFoundation model was trained on a custom web-scraped dataset of 50M single cells. Based on CELLxGENE data, there looks to be a linear relationship between tokens and cell count. From this we can estimate that 50M cells correspond to roughly 93B tokens. Let’s round up to 100B, which is close to the available data on CELLxGENE.
Using the Chinchilla formula, we can assess predicted loss across different model sizes and data scales. Essentially, this allows us to explore how various model sizes would perform with different data-constraints. At the current data regime, it would make sense to train a model at least an order of magnitude larger scale, probably close to 5B. This checks out with the 20:1 tokens to parameter rule of thumb. After that, you need more data to trade off diminishing returns.
Publicly available data might be able to provide this earlier than I personally expected. In the optimistic scenario you are at 500B tokens before 2027 🧐… One interpretation is that without proprietary assay technology, private companies may not truly possess a “data-platform advantage” in such a race to collect data. It makes sense that publicly available data is increasing more rapidly than I expected: crowd-sourcing parallelizes data collection and distributes cost.
Caveats
It's important to reiterate: there is no evidence to support the assumption that the scaling laws observed in NLP translate directly to scRNA-seq data. While some core computational processes in transformer-based models may be similar, the modalities are vastly different. Each new domain of information requires comprehensive scaling experiments to determine its specific characteristics. And, frankly, this scaling law work is “old” in the fast-moving field of AI. Findings from just a few years ago may be upended as we discover how to dynamically distribute compute.6
Additionally, it remains unclear whether advancements in pretraining will lead to tangible improvements in downstream tasks related to drug discovery. By the way, what are these critical tasks? Few have been comprehensively benchmarked and defined. Numerous unanswered questions remain.
I plan to conduct similar analyses on sequences and structures to address broader data availability questions. If you're interested in collaborating on these explorations, feel free to reach out!
https://www.cs.utexas.edu/~eunsol/courses/data/bitter_lesson.pdf\
https://arxiv.org/abs/1810.04805
ESM: https://www.pnas.org/doi/10.1073/pnas.2016239118
NT: https://www.biorxiv.org/content/10.1101/2023.01.11.523679v3
Geneformer: https://www.nature.com/articles/s41586-023-06139-9
https://arxiv.org/abs/2001.08361
https://arxiv.org/abs/2203.15556
https://arxiv.org/abs/2404.02258