The Central Dogma
When we think of modeling a system, it is often beneficial to decompose the system into constituent parts and formalize how these parts interact. How would you apply this principle to a living system? You might call upon The Central Dogma of molecular biology (TCD) to scaffold a solution. Contrary to popular understanding, TCD is not simply DNA → RNA → protein. Here’s the original definition by Francis Crick:
The Central Dogma. This states that once "information" has passed into protein it cannot get out again. In more detail, the transfer of information from nucleic acid to nucleic acid, or from nucleic acid to protein may be possible, but transfer from protein to protein, or from protein to nucleic acid is impossible. Information here means the precise determination of sequence, either of bases in the nucleic acid or of amino acid residues in the protein.
While the transfer of information from DNA to RNA and from RNA to protein are essential components of TCD, we also observe DNA → DNA and RNA → RNA (replication) and even RNA → DNA (reverse transcription). Crick's original postulate encompasses these various methods of information transfer.
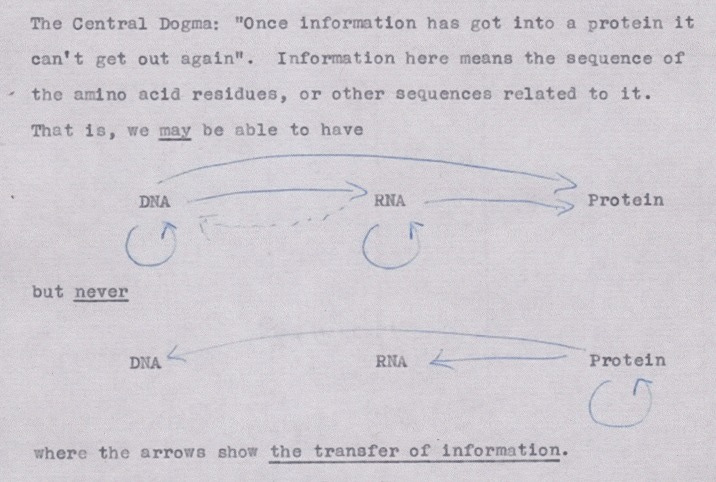
In essence, TCD defines the bounds of information transfer in molecular biology. While the core principles of TCD hold true, numerous methods of information transfer have been discovered at the molecular level. Examples include RNA splicing, regulation through non-coding RNA, epigenetic regulation, and even protein structural alteration mediated by prions.
It appears straightforward to model biology through this framework. However, from a computational perspective, this framework overlooks crucial aspects of the information feedback systems that make biology dynamic. Biology is not static; it operates (1) over time and (2) in response to the environment.
I believe that many of our modeling frameworks are biased by the available measurement methods. Currently, experimental methods for densely sampling biological systems across either of these variables are rudimentary.
Case study: the lac operon
How does information transfer back from protein to DNA? This process does not occur through transfer of sequence information, but rather is manifested directly through biophysical phenomena. This is what makes biology dynamic and responsive to internal and external variables over time. Picture the cell as a "soup" of various molecules, each with distinct chemical properties and concentrations. These characteristics influence the likelihood of interactions between any two molecules. Through time, these biophysical interactions coalesce into complex systems with intertwined feedback loops, shaping the cell's behavior.
The lac operon is a well-known case study of regulatory dynamics in prokaryotic systems. This operon controls the metabolism of lactose. When glucose is scarce and lactose is present, the lac operon is activated.

The lac repressor protein, endogenously expressed, binds to DNA, preventing the expression of the genes necessary for lactose metabolism. The biophysical compatibility between the protein and this specific DNA sequence results in the formation of a stable binding complex. When lactose enters the cell from the environment, it binds to the lac repressor protein, causing a conformational (structural) change that releases the repressor from the DNA. This allows RNA polymerase to bind and transcribe the genes.
Another condition must be met for the system to fully activate: glucose must be absent. This condition ensures that the cell only metabolizes lactose when glucose is not available.
We will leave it there, but the complexity of this system quickly compounds as you realize there is really no end to this self-contained system. These principles manifest the observable organisms we are.
The bioML stack
Applying a computational perspective has shifted my understanding of biological systems beyond the traditional focus on DNA, RNA, and protein. Three modalities characterize my bioML stack: sequence, structure, and systems. Each of these information-carrying modalities exhibit varying levels of expressivity.
Sequence:
The linear arrangement of nucleotides in DNA or amino acids in proteins.
Nature: Static information (storage).
Sequences are relatively stable and remain largely unchanged over time.
They provide the blueprint for the structure and function of biomolecules.
Examples:
DNA sequences store genetic information, ultimately determining an organism's primary traits.
Protein sequences define structure and function in cellular processes.
Structure:
The three-dimensional organization of biomolecules like proteins and nucleic acids.
Nature: Semi-dynamic information.
Structures undergo conformational changes, but they remain within a defined range.
These changes influence the molecule's functional properties.
Examples:
Protein structures determine binding sites, catalytic activity, etc.
Nucleic acid structures affect stability, replication, and transcription.
Systems (concentration):
“Systems” really means “concentration” or “abundance” but it is nice to preserve the alliteration. Encompasses dynamic changes in the quantity and composition of biomolecules within a cell or organism.
Nature: Dynamic information.
Systems experience continuous fluctuations in the abundance of different molecules.
These changes respond to cellular needs, environmental factors, and regulatory mechanisms.
Examples:
Gene regulation: mRNA and protein abundance varies, controlling gene expression.
Metabolic pathways: Metabolites and enzyme abundance fluctuates, regulating metabolism.
This stack is far from any “information theoretic” formalism. But this fuzzy categorization helps me frame biological systems in terms of their constituent parts. These components can be independent, dependent, interdependent or composed in a variety of ways. Consider the lac operon as an example, which encompasses instances of each of these modalities.
The core challenge of modeling biological systems is approximating how different modalities interact to drive a phenotype. We have good methods to measure these modalities. Learned models attempt to capture what happens between two measured states. At an extreme level, you can think of this as backpropagating from a phenotype all the way back down to the DNA sequence.
However, this simplification overlooks two crucial variables: time and environment. Biological systems function over time and are influenced by information intervention and feedback. Only under ideal conditions, such as a perfect 72-degree day in the spring, might all systems be in homeostasis. Furthermore, the environment not only affects organisms but also vice versa. Consider glucose metabolism: glucose is available, and then not. Humans provide numerous examples of altering the environment to meet our biological needs.
This illustrates the complexity of biological systems. We operate within a balance of interactions where it is often difficult to discern where one system ends and another begins.





Awesome post Will! You might like the work that stems from Karl Friston’s Free Energy Principle (especially it’s specialization for living systems called “active inference”). The goal is to formularize living systems as agents operating over time that are interacting with other agents and their environment (which also is an agent), with the goal of processing information in order to minimize sensory surprise. Sounds like there might be some similarities between your perspective and this communities’ perspective!